Impacts of Boundary Conforming Meshes on Electrical Cardiac Simulation
|
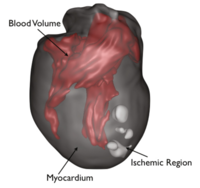
Computational simulation has become an indispensable tool in the study of both basic mechanisms and pathophysiology of all forms of cardiac electrical activity. Such simulations depend heavily on geometric models that are either realistic or even patient specific. These models consist of a connected mesh of sometimes millions of polygonal elements that must capture the complex external shapes and internal boundaries among regions of the heart. The resulting meshes can be non-conforming, i.e., they have element faces that fail to align with the tangents of the surfaces or boundaries and consequently the elements are a poor approximation of these smooth surfaces and boundaries. We hypothesize that such jagged, non-conforming meshes, which are often preferred, as they are easier to create, produce local artifactual concentrations of current that lead to simulation errors large enough to distort the resulting potential fields and generate misleading results. We tested this hypothesis on two types of numerical approximation used in bioelectric simulations: bidomain, and reaction-diffusion bidomain. Comparison with gold standard results for the monodomain and bidomain simulations showed that errors within a few elements (3-5) of the surface could be as large as 10-32%. The root mean squared error over the entire mesh was more modest, ranging from 1-6%. In the case of reaction diffusion simulations, by contrast, such meshing errors accounted for only an insignificant component of overall simulation uncertainty. These findings lead to the conclusion that while non-conforming meshes are certainly less costly to produce, their use can result in substantial local errors that depend highly on the specific problem of interest and the numerical approximation approach. |
[DOI/EE link]
@inproceedings{SLTWM12,
address = {San Jose, CA},
author = {Darrell J. Swenson and Joshua A. Levine and Jess D. Tate and Ross T. Whitaker and Rob S. MacLeod},
booktitle = {21st International Meshing Roundtable},
ee = {http://dx.doi.org/10.1007/978-3-642-33573-0_34},
month = {10},
pages = {585--602},
publisher = {Springer},
title = {Impacts of Boundary Conforming Meshes on Electrical Cardiac Simulation},
year = {2012}
}